Difference between revisions of "Profiling GEOS-Chem with the TAU performance system"
(→Installing TAU) |
|||
| (26 intermediate revisions by one other user not shown) | |||
| Line 1: | Line 1: | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
== Overview == | == Overview == | ||
| Line 14: | Line 5: | ||
== Installing TAU == | == Installing TAU == | ||
| − | The best way to | + | The best way to build TAU is with Spack. |
| − | + | == Compiling and running GEOS-Chem with TAU == | |
| − | + | You can use the <tt>tau_exec</tt> command to profile GEOS-Chem in order to reveal computational bottlenecks. Below is a job script that you can use. | |
| − | + | NOTE: The <tt>gcc.env</tt> file is the [https://github.com/geoschem/geos-chem/issues/637#issuecomment-788968697 script that loads your Spack modules]. | |
| − | + | ||
| − | + | #!/bin/bash | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | # | + | #SBATCH -c 24 |
| − | + | #SBATCH -n 1 | |
| + | #SBATCH -p YOUR-QUEUE-NAME | ||
| + | #SBATCH --mem=15000 | ||
| − | # | + | ### geoschem.profile.run |
| − | + | ### Job script to run GEOS-Chem with TAU profiling | |
| + | |||
| + | source ~/.bashrc # Load bash settings | ||
| + | source ~/gcc.env # Environment file that loads TAU module from Spack | ||
| − | # | + | # Set the # of OpenMP threads to the # of cores we requested |
| − | export | + | export OMP_NUM_THREADS=24 |
| − | + | # Run GEOS-Chem Classic and have TAU profile it | |
| + | srun -c $OMP_NUM_THREADS tau_exec -T serial,openmp -ebs -ebs_resolution=function ./gcclassic > log 2>&1 | ||
| − | + | If you use SLURM, you can submit this with | |
| − | + | sbatch geoschem.profile.run | |
| − | + | Otherwise you can just run it interactively: | |
| − | + | ./geoschem.profile.run | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
== Using ParaProf to create plots from the profiling data == | == Using ParaProf to create plots from the profiling data == | ||
| Line 104: | Line 59: | ||
To save the plot, select "Save as Bitmap Image" from the File menu. In the Save Image File window, you may select the output type (JPEG File or PNG file) and specify the file name and location. | To save the plot, select "Save as Bitmap Image" from the File menu. In the Save Image File window, you may select the output type (JPEG File or PNG file) and specify the file name and location. | ||
| − | |||
| − | |||
Latest revision as of 18:12, 24 October 2023
Contents
Overview
The TAU Performance System is a profiling tool for performance analysis of parallel programs in Fortran, C, C++, Java, and Python. TAU uses a visualization tool, ParaProf, to create graphical displays of the performance analysis results.
Installing TAU
The best way to build TAU is with Spack.
Compiling and running GEOS-Chem with TAU
You can use the tau_exec command to profile GEOS-Chem in order to reveal computational bottlenecks. Below is a job script that you can use.
NOTE: The gcc.env file is the script that loads your Spack modules.
#!/bin/bash #SBATCH -c 24 #SBATCH -n 1 #SBATCH -p YOUR-QUEUE-NAME #SBATCH --mem=15000 ### geoschem.profile.run ### Job script to run GEOS-Chem with TAU profiling source ~/.bashrc # Load bash settings source ~/gcc.env # Environment file that loads TAU module from Spack # Set the # of OpenMP threads to the # of cores we requested export OMP_NUM_THREADS=24 # Run GEOS-Chem Classic and have TAU profile it srun -c $OMP_NUM_THREADS tau_exec -T serial,openmp -ebs -ebs_resolution=function ./gcclassic > log 2>&1
If you use SLURM, you can submit this with
sbatch geoschem.profile.run
Otherwise you can just run it interactively:
./geoschem.profile.run
Using ParaProf to create plots from the profiling data
In your run directory, there should be one or more profile.* files. The number of profile.* files will depend on the number of CPUs that you use for your GEOS-Chem simulation. To pack all of the profiling data into a single file, type:
paraprof --pack GEOS-Chem_Profile_Results.ppk
Then run paraprof on the packed format (.ppk) file using:
paraprof GEOS-Chem_Profile_Results.ppk
This will open two windows, the ParaProf Manager window and the Main Data window. For more information on how to interpret the profiling data, see the ParaProf User's Manual.
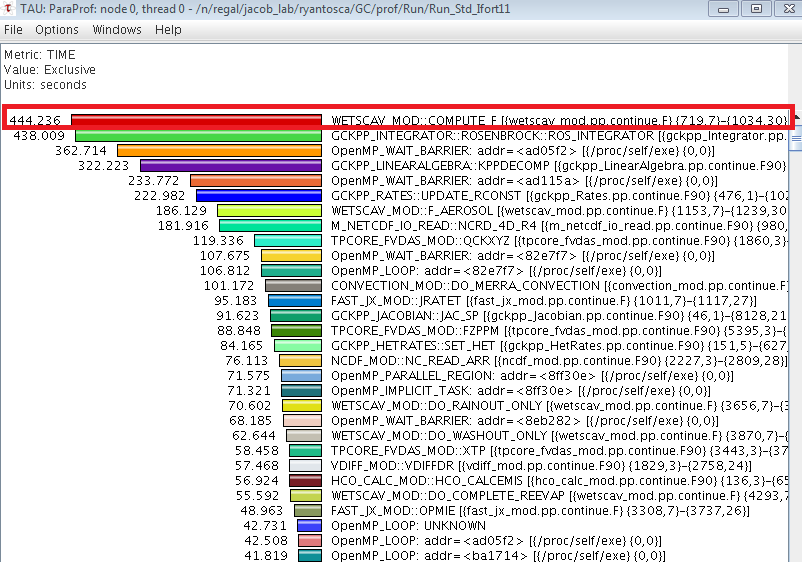
If you click on the the bar labeled "thread0" in the ParaProf manager window, you can generate a plot that looks like this:
The value displayed, the units, and the sort order can be changed from the Options menu. The time that each subroutine spent on the master thread is displayed as a histogram. By examining the histogram you can see which routines are taking the longest to execute. For example, the above plot shows that the COMPUTE_F routine (highlighted with the red box) is spending 444 seconds on the master thread, which is longer than the Rosenbrock solver takes to run. This is a computational bottleneck, which was ultimately caused by an unparallelized DO loop.
To save the plot, select "Save as Bitmap Image" from the File menu. In the Save Image File window, you may select the output type (JPEG File or PNG file) and specify the file name and location.