Process Analysis Diagnostics
Process Analysis Diagnostics
Contents
Overview - What is Process Analysis?
Process-based Analysis examines the change in each species due to each process and reaction. Models predict atmospheric state, which in a time-series can be used to create net-change of each species. What this cannot tell us, is which processes led to that change. To supplement state (or concentration, GEOS-Chem has long archived emissions and employed advanced diagnostics to predict gross chemical production or loss. Process Analysis goes a step further archiving grid-cell budgets for each species, and decomposing gross production/loss into individual reaction contributions. Process Analysis extensions are currently available in [CAMx], [WRF-Chem], [CMAQ], and now GEOS-Chem. This allows for direct comparisons of models at a fundamental, process level.
Details - No really, what is it?
The budget is built from process contributions and reaction throughput. Process contributions are referred to as Integrated Process Rates (IPR) and can be calculated because of operator splitting. Each process is distinctly called, and state before and after the call can be differenced to see the process contribution. The process contributions (ΔC) are conceptually shown in the figure above.
Reaction contributions are more complex to calculate than IPR, but essentially expand the chemistry process ΔCchem. Reaction contributions are constructed from integrated reaction rates (IRR). IRR are calculated according to equation 1. The rate coefficient (k) is evaluated by calcrate.f and then SMVGEAR or KPP solve over short timesteps (<math>t_s</math>). To calculate the IRR, we must add code to SMVGEAR or KPP that finds successful steps and archives equation 1.
- IRRrxn = Δ ts· krxn · ∏rct[Crct]
- ΔCi,rxn = αi,rxnIRRrxn
- ΔCi,chem = Σrxn ∈ rxnsΔCi,rxn
Development Status - What features are available?
Integrated Process Rates
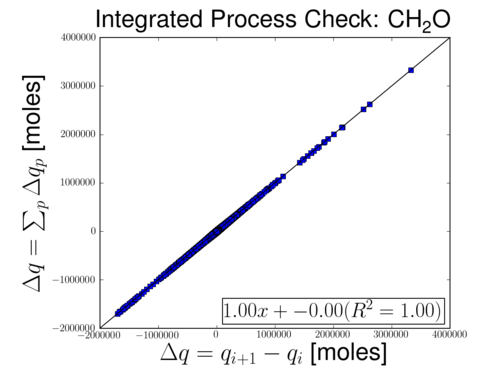
Process rates are fully functional, but will be extended in the future. Right now, the process are able to perfectly account for the change in concentrations. However, some processes are lumped and others contain directional information that is not currently captured.
- Transport
- Wet deposition
- Heterogeneous Chemistry
- Gas-phase Chemistry
- Convective Mixing
- Emissions*
- Dry deposition*
*Gas-phase emissions and dry deposition are solved either in the chemical solver or in asymmetric convection routines. Separation of processes within convection has not yet been implemented.
The figure below shows change in concentration on the x-axis and sum of processes on the y-axis.
Integrated Reaction Rates
Integrated Reaction Rates are fully functional with two solver options, but more QA is needed.
- Sparse Matrix Vector Gear: available
- Kinetic Pre-Processor
- Rosenbrock (coming soon)
- LSODES (available upon request)
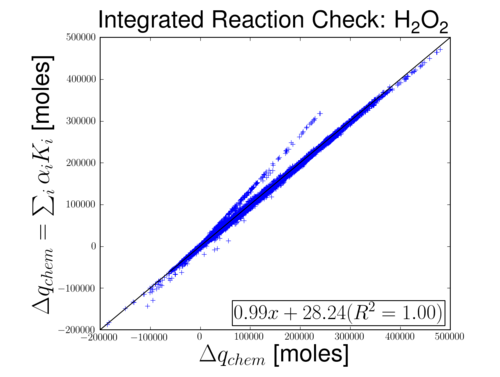
The image below shows the chemistry process on the x-axis and the sum of reaction contributions. Currently, there is a discrepancy for some grid cells. It may be related to the species level error correction in the SMVGEAR solver.
A KPP LSODES solver with IRR is available, but GEOS-Chem uses Rosenbrock by default. Efforts have been made to implement IRR in Rosenbrock, but more work is necessary to account for the 4-stage solution contributions.
To do:
- Incorporate error correction if necessary
- Complete KPP Rosenbrock solver implementation
Installation and Application - How can I use it?
The code is available in a github repository that is currently available upon request from Barron Henderson at the University of Florida.
After obtaining the code, compiling it is the same as compiling standard GEOS-Chem. At some point in the future, a windowing interface should be added to input.geos. Right now, the spatial domain is hard coded in the pa_mod.F file. Edit pa_mod.F as necessary and then compile.
After compiling the code, running it is identical to running standard GEOS-Chem. In addition to the ctm.bpch file, ctm.pa.YYYYMMDDHH and ctm.irr.YYYYMMDDHH files will be output for each day. Post-processing the results can be done by making minor modifications to the diaginfo.dat and tracerinfo.dat files. With the distributed code, the modifications are available. This too should be integrated into the standard approach.
A common analysis approach is to integrate within the PBL and within some spatial region. Tools to accomplish this are available through the Python-based Process Analysis project.
Working with integrated reaction rates requires applying stoichiometry to the rates outside of the model. This can be accomplished easily with the Python-based Environment for Reaction Mechanisms/Mathematics project.