Profiling GEOS-Chem with the TAU performance system
Previous | Next | Guide to GEOS-Chem performance
- Parallelizing GEOS-Chem
- GEOS-Chem 7-day timing tests
- GEOS-Chem scalability
- GEOS-Chem 1-month benchmark timing results
- Profiling GEOS-Chem with the TAU performance system
- Speeding up GEOS-Chem
NOTE: This page is still under development. The Spack installation guide is still being validated.
Contents
Overview
The TAU Performance System is a profiling tool for performance analysis of parallel programs in Fortran, C, C++, Java, and Python. TAU uses a visualization tool, ParaProf, to create graphical displays of the performance analysis results.
Installing TAU
NOTE: These instructions are still in flux.
The best way to install TAU is to use Spack.
(1) Download Spack as described at |Github issue geoschem/geos-chem #637].
(2) Define environment variables for Spack as described HERE.
(3) Check if Spack can find your compiler as described HERE.
(4) Install TAU with the following Spack command:
spack install tau@2.28.1 openmp=True
You can also force installation with a specific compiler version by typing:
spack install tau@2.28.1 %gcc@8.2.0 openmp=True
(5) Define environment variables for TAU as shown below.
# TAU home location export TAU_HOME=$(spack location -i tau) # Put the location of tau_f90.sh in the search path export PATH=$PATH:$TAU_HOME/bin # Define options for TAU export TAU_OPTIONS="-optRevert -optVerbose -optPreProcess -optContinueBeforeOMP -optPdtGnuFortranParser" # Define location of TAU makefile export TAU_MAKEFILE=$TAU_HOME/x86_64/lib/Makefile.tau-papi-pthread-pdt-openmp
You can place these into one of your system environment files so that they are executed on startup.
Compiling and running GEOS-Chem with TAU
To profile GEOS-Chem with TAU, you must first compile with the TAU_PROF=y Makefile option, e.g.:
# Remove files from a previous compilation with TAU (if necessary) make tauclean # Compile with TAU profiling make TAU_PROF=y ...etc. other makefile options ...
It is important that you compile on a single processor (i.e. don't pass -j4 or -j8) to allow TAU to properly instrument the code.
The TAU_PROF=y option will set COMPILE_CMD :=tau_f90.sh instead of COMPILE_CMD :=$(FC) where FC is gfortran or ifort.
Once you have compiled GEOS-Chem with TAU_PROF=y, you can run GEOS-Chem as you normally would. GEOS-Chem will create profile.* files containing the profiling information.
Using ParaProf to create plots from the profiling data
In your run directory, there should be one or more profile.* files. The number of profile.* files will depend on the number of CPUs that you use for your GEOS-Chem simulation. To pack all of the profiling data into a single file, type:
paraprof --pack GEOS-Chem_Profile_Results.ppk
Then run paraprof on the packed format (.ppk) file using:
paraprof GEOS-Chem_Profile_Results.ppk
This will open two windows, the ParaProf Manager window and the Main Data window. For more information on how to interpret the profiling data, see the ParaProf User's Manual.
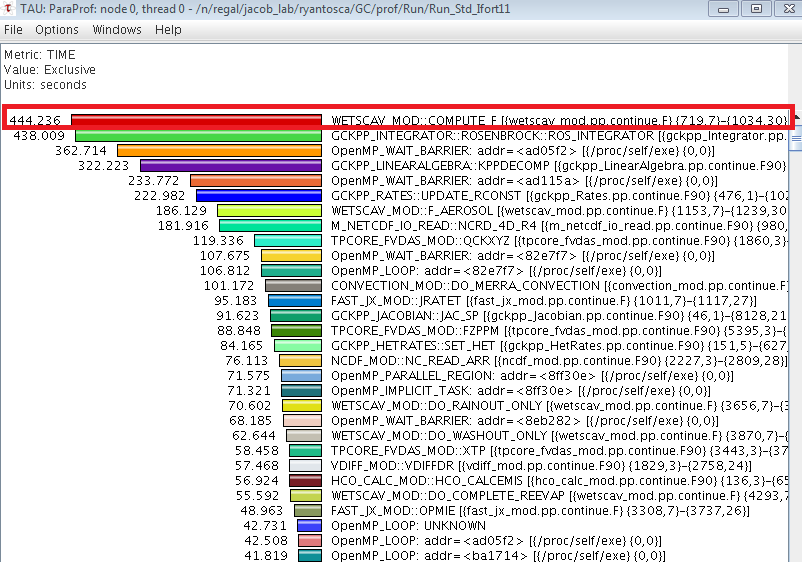
If you click on the the bar labeled "thread0" in the ParaProf manager window, you can generate a plot that looks like this:
The value displayed, the units, and the sort order can be changed from the Options menu. The time that each subroutine spent on the master thread is displayed as a histogram. By examining the histogram you can see which routines are taking the longest to execute. For example, the above plot shows that the COMPUTE_F routine (highlighted with the red box) is spending 444 seconds on the master thread, which is longer than the Rosenbrock solver takes to run. This is a computational bottleneck, which was ultimately caused by an unparallelized DO loop.
To save the plot, select "Save as Bitmap Image" from the File menu. In the Save Image File window, you may select the output type (JPEG File or PNG file) and specify the file name and location.
--Melissa Sulprizio (talk) 22:48, 8 February 2017 (UTC)
--Bob Yantosca (talk) 16:55, 10 February 2017 (UTC)