Common GEOS-Chem error messages
NOTE: Some of the posts below contain obsolete information. We shall leave these here for your convenience, but have marked them as obsolete.
Here is a list of some commonly-encountered GEOS-Chem error messages. Also be sure to visit our Machine issues and portability wiki page for a list of compiler-specific issues.
Contents
- 1 Compile-time warnings and errors
- 2 Compile-time errors caused by compiler bugs
- 3 Run-time crashes and abnormal exits
- 4 Segmentation faults
- 5 Other memory-related errors
- 6 Obsolete error messages
- 6.1 Error caused by MEGAN biogenic emissions
- 6.2 I/O Error #29
- 6.3 Problem reading binary punch file
- 6.4 Error when reading the "restart_gprod_aprod" file
- 6.5 File ann_mean_trop.geos5.* not found
- 6.6 Problem reading GEOS-4 TROPP files
- 6.7 OPEN failure
- 6.8 SMVGEAR "Too many decreases of YFAC" error
- 6.9 JXTRA error in FAST-JX photolysis
- 6.10 Too many levels in photolysis code
- 6.11 Error computing F_OF_PBL
- 6.12 LISOPOH error
- 6.13 Fatal error in IFORT
- 6.14 A3 met fields not found
Compile-time warnings and errors
In this section we discuss some compilation warnings that you may encounter.
- Warnings are not generally fatal—GEOS-Chem will usually continue to compile while an informational message is displayed.
- Errors, on the other hand, will halt the GEOS-Chem compilation process.
Error in opening the compiled module file
Example:
RnPbBe_mod.F(118): error #7002: Error in opening the compiled module file. Check INCLUDE paths. [CHEMGRID_MOD]
USE CHEMGRID_MOD, ONLY : ITS_IN_THE_STRATMESO
----------^
RnPbBe_mod.F(118): error #6580: Name in only-list does not exist. [ITS_IN_THE_STRATMESO]
USE CHEMGRID_MOD, ONLY : ITS_IN_THE_STRATMESO
-------------------------------------^
compilation aborted for RnPbBe_mod.F (code 1)
This error message usually indicates indicates a typo in the Makefile dependencies section. For example, the above error was caused by this Makefile code
rnpbbe_mod.o : RnPbBe_mod.F \ chemgrid_mod.o diag_mod.o \ hco_interface_mod.o
because the capitalization of the object file name did not exactly correspond to the source code file name. The compilation process died with the above error because it was expecting to find a module file named RnPbBe_mod.o but could not.
--Bob Yantosca (talk) 16:19, 27 February 2017 (UTC)
Module file cannot be read
If you should encounter this type of error:
ifort -cpp -w -O2 -auto -noalign -convert big_endian -openmp -Dmultitask -c time_mod.f
fortcom: Error: time_mod.f, line 259: This module file was generated for a different
platform or by an incompatible compiler or compiler release. It cannot be read. [JULDAY_MOD]
USE JULDAY_MOD, ONLY : JULDAY, CALDATE
Then this means that you are trying to link to previously-created *.mod files that were generated by a different compiler. Making clean and re-compiling from scratch should solve this problem.
--Bob Y. 13:39, 1 July 2008 (EDT)
Internal threshold was exceeded
This warning is specific to the Intel Fortran Compiler. It usually happens when you try to optimize a complex module or subroutine. Please see this post on the software.intel.com site for a full explanation.
--Bob Y. 15:32, 22 August 2012 (EDT)
Compile-time errors caused by compiler bugs
A few GEOS-Chem errors have been traced to bugs in the compiler that was used to build the GEOS-Chem executable. For your convenience, we have collated a list of these issues. Please see our our Known issues caused by compiler bugs wiki page for more information.
--Bob Yantosca (talk) 19:13, 13 April 2016 (UTC)
Failed in XMAP_R4R4 error
If you are using the Intel Fortran Compiler 15, then you may encounter an error such as this:
forrtl: severe (408): fort: (2): Subscript #1 of the array LON2 has value 1 which is greater than the upper bound of -1 Image PC Routine Line Source libifcoremt.so.5 00002B9EFA2188D3 Unknown Unknown Unknown geos.mp 00000000011FCE35 regrid_a2a_mod_mp 1914 regrid_a2a_mod.F90 libiomp5.so 00002B9EFB70A8A3 Unknown Unknown Unknown
Cause: A compiler bug in Intel Fortran Compiler version 15.
Solution: If you are using array-out-of-bounds checking, make sure to compile GEOS-Chem with these flags: BOUNDS=y DEBUG=y. For more information, see this post on our HEMCO wiki page.
--Bob Yantosca (talk) 17:18, 25 January 2016 (UTC)
Seg fault in HEMCO with Intel Fortran 12, 13, or 14
In GEOS-Chem versions from v10-01 up to v11-01j, this error can occur with versions 12, 13, and 14 of the Intel Fortran Compiler.
forrtl: severe (174): SIGSEGV, segmentation fault occurred Image PC Routine Line Source ... geos 0000000000ADDCF4 hcoio_dataread_mo 2810 hcoio_dataread_mod.F90 geos 0000000000AD84DE hcoio_dataread_mo 2499 hcoio_dataread_mod.F90 geos 0000000000AB8971 hco_readlist_mod_ 438 hco_readlist_mod.F90 geos 0000000000AB8398 hco_readlist_mod_ 267 hco_readlist_mod.F90 geos 0000000000AA99A6 hco_driver_mod_mp 138 hco_driver_mod.F90 geos 0000000000844C07 hcoi_gc_main_mod_ 546 hcoi_gc_main_mod.F90 geos 0000000000801A2E emissions_mod_mp_ 172 emissions_mod.F90 geos 00000000006720B7 MAIN__ 834 main.F
For a workaround, please see this post on our HEMCO wiki page.
NOTE: A similar workaround has been introduced into the v11-01 public release, which uses a newer version of HEMCO.
--Bob Yantosca (talk) 21:50, 6 January 2017 (UTC)
Run-time crashes and abnormal exits
In this section, we provide information about some commonly-reported run-time errors that cause GEOS-Chem to halt executing.
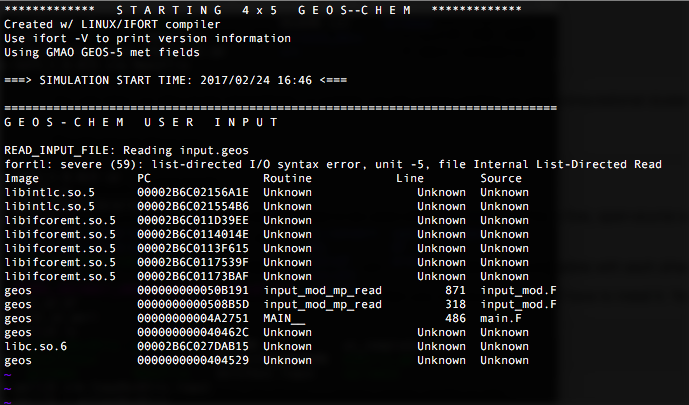
List-directed I/O syntax error
The above error message indicates that the simulation crashed at line 871 in GeosCore/input_mod.F. This means there was an issue while reading the input.geos file (located in the run directory). For example, GEOS-Chem might have expected numeric input, but instead a character was read from input.geos, thus causing a read error. This type of error can occur the input.geos corresponds to a version of GEOS-Chem that is different from yours.
This error is not limited to input.geos; it can happen for any text file that is being read from disk (both in GEOS-Chem and in any other Fortran programs you may write).
--Bob Yantosca (talk) 15:51, 27 February 2017 (UTC)
No output scheduled on last day of run
If you encounter this error at the start of your GEOS-Chem simulation:
==========================================================================
GEOS-CHEM ERROR: No output scheduled on last day of run!
STOP at IS_LAST_DAY_GOOD ("input_mod.f")
==========================================================================
This means that you have not told GEOS-Chem to save out diagnostic data on the day that your simulation ends. GEOS-Chem adds this error check in order to prevent you from running a long simulation only to have no diagnostics printed out at the end of the run.
For more information on how to schedule diagnostic output in GEOS-Chem, please see the section entitled ""the OUTPUT MENU section of the input.geos file" in our GEOS-Chem Online Users' Guide.
--Bob Y. 12:56, 15 July 2011 (EDT)
HEMCO Run Error
Errors messages containing "HCO" originate in the HEMCO emissions component. For example:
=============================================================================== GEOS-CHEM ERROR: HCO_RUN STOP at HCOI_GC_RUN (hcoi_gc_main_mod.F90) ===============================================================================
Additional helpful diagnostic information can be found in the HEMCO log file, which is usually named HEMCO.log.
--Chris Holmes (talk) 15:42, 24 June 2015 (UTC)
Updated error message for v11-01
In GEOS-Chem v11-01 and higher versions, additional text instructs the user to also check the HEMCO log file.
===============================================================================
GEOS-CHEM ERROR: HCO_RUN
HEMCO ERROR: Please check the HEMCO log file for error messages!
STOP at HCOI_GC_RUN (GeosCore/hcoi_gc_main_mod.F90)
===============================================================================
--Bob Yantosca (talk) 20:50, 6 January 2017 (UTC)
Allocation error
If your GEOS-Chem simulation dies with this error output:
=============================================================================== GEOS-CHEM ERROR: Allocation error in array: MY_ARRAY STOP at alloc_err.f ===============================================================================
then this means that you do not have enough memory to run your simulation. This type of error can frequently occur if you are running a full-chemistry simulation at the 2° x 2.5° global grid, or one of the 0.5° x 0.666° nested grids. You may need to tell your queuing system to make more memory available for your GEOS-Chem simulation. Ask your IT staff for details.
--Bob Yantosca (talk) 22:15, 6 January 2017 (UTC)
Permission denied error
If you receive this error:
v11-01.run: Permission denied.
after having submitted a run script to a queue system (such as SLURM or Grid Engine), then doublecheck the Unix permissions of your v11-01.run script. If the script does not have the Unix "execute" permission then the queue system will not be able to run it.
Use the Unix chmod command to make your script executable
chmod 755 v11-01.run
and then re-submit the script to the queue system.
--Bob Yantosca (talk) 22:17, 6 January 2017 (UTC)
N_/L2_/L2-cutoff JXTRA error
---> DATE: 2013/11/13 GMT: 19:00 X-HRS: 307.000
- Found all 44 GEOS-FP A1 met fields for 2013/11/13 19:30
N_/L2_/L2-cutoff JXTRA: 601 96 0.00
N_/L2_/L2-cutoff JXTRA: 601 96 0.00
... run dies later with NaN's in chemistry ...
If you are using a version of GEOS-Chem prior to v11-01d, then you might have experienced this error, where a simulation using GEOS-FP or MERRA met fields dies in the chemistry solver after many such "JXTRA" warnings are printed to the log file.
This error has been attributed to a bug in the GEOS-FP and MERRA convection module. This issue was resolved in v11-01d. Upgrading to the GEOS-Chem v11-01 public release code or (patching your GEOS-Chem v10-01 code with this fix) will solve the issue.
--Bob Yantosca (talk) 23:12, 27 February 2017 (UTC)
KPP "Step size too small" error
The following abnormal exit from the KPP chemical solver:
Forced exit from Rosenbrock due to the following error:
--> Step size too small: T + 10*H = T or H < Roundoff
T= 3044.21151383269 and H= 1.281206877135470E-012
...
1
Forced exit from Rosenbrock due to the following error:
--> Step size too small: T + 10*H = T or H < Roundoff
T= 3044.21151383269 and H= 1.281206877135470E-012
failed twice !!!
indicates that the chemistry could not converge to a solution in the given grid box. Possible reasons for this could be:
- A particular tracer has numerically underflowed or overflowed. This can happen especially in the aerosol chemistry and equilibrium routines, where many exponentials and logarithms are used in the algorithms.
- The restart file is not appropriate for the given simulation. For example, if the restart file was created using the Synoz O3 flux boundary condtion, but you have turned on the Linoz stratospheric O3 chemistry, then this mismatch can cause the solver not to converge. You can try switching to a restart file generated from a simulation with the same input options as the simulation that you wish to perform.
You may have to manually adjust the convergence criteria in the GEOS-Chem code to fix this condition.
--Bob Y. 11:30, 9 November 2010 (EST)
Negative tracer found in WETDEP
If your simulation encounters negative (or NaN) tracer concentrations in the WETDEP routine, then this can be an indication of a problem further upsteam, perhaps in the aerosol routines (highly probable if the tracer is SO4, SO4s, HNO3, SO2, or NH3). We have fixed some of these bugs by making the code more robust. If you are using a GEOS-Chem version prior to v8-01-01, then you should get ftp://ftp.as.harvard.edu/pub/geos-chem/patches/v8-01-01/ these patches. (These patches have been added to the standard GEOS-Chem code in versions higher than v8-01-01.) Please see the following links for more information:
- Values of F_PRIME > 1 in routine WETDEP
- Negative tracer due to negative RH values in the met field data
- Negative tracer in routine WETDEP
- Negative tracer in routine WETDEP #2
- A bug in RPMARES was also leading to a crash in WETDEP.
If the fixes above do not solve your problem, you will need to debug. The first step is to use few calls to CHECK_STT (from tracer_mod.f) to isolate the part of the code where negative tracers are created. This can be done quite fast if the code dies early enough in the run.
--Bob Y. 12:50, 15 July 2011 (EDT)
Segmentation faults
If your simulation dies with a segmentation fault error, this means that GEOS-Chem tried to access an invalid memory location. We list several instances of segmentation faults below.
Severe(174) SIGSEGV error
NOTE: In this section, we shall use the IFORT compiler error messages. You may get a slightly different error message if you are using a different compiler (such as PGI).
If you compiled GEOS-Chem with the IFORT compiler, you may encounter the following error message:
forrtl: severe (174): SIGSEGV, segmentation fault occurred
This means that a segmentation fault (i.e. memory error) has occurred during your GEOS-Chem simulation. This can be caused by:
Array-out-of-bounds error
Most often, a segmentation fault indicates an array out-of-bounds condition. To find out more information about where this error is occurring, recompile GEOS-Chem with the following Makefile options:
make realclean make BOUNDS=yes TRACEBACK=yes
The BOUNDS=yes option will turn on Array Out-of-Bounds error checking. The TRACEBACK=yes option will print out the Error Stack, or a list of routines that were called, and the line at which the error occurred. These options will provide more detailed error output.
NOTE: TRACEBACK=yes is turned on by default in v11-01 and higher versions.
After recompiling, you should receive an error message such as:
forrtl: severe (408): fort: (3): Subscript #1 of the array PBL_THICK has value -1000000 which is less than the lower bound of 1
This tells you that there is a problem with a certain array. Use the Unix grep command to search for all instances of this array in the GEOS-Chem source code:
grep -i PBL_THICK *.f*
and search for the problem.
NOTE: In the above example, we manually forced an out-of-bounds error with this line of code:
!### FORCE OOB error for testing
PBL_THICK(-1000000,J) = BLTHIK
Removing this line will fix the error.
--Bob Yantosca (talk) 22:19, 6 January 2017 (UTC)
Invalid memory access
A segmentation fault can also happen if GEOS-Chem makes an reference to a memory location that is invalid. You may see an error message such as this:
severe (174): SIGSEGV, segmentation fault occurred This message indicates that the program attempted an invalid memory reference. Check the program for possible errors.
This can happen if you are trying to read data from a file into an array, but the array is too small to hold all of the data. You can use a debugger (such as Totalview or IDB) to try to diagnose the situation. You may receive an error message from the debugger similar to this one:
Thread received signal SEGV stopped at [<opaque> for_read_seq_xmit(...) 0x40000000006b6500] Information: An <opaque> type was presented during execution of the previous command. For complete type information on this symbol, recompilation of the program will be necessary. Consult the compiler man pages for details on producing full symbol table information using the '-g' (and '-gall' for cxx) flags.
Usually, increasing the size of the array (i.e. until it is large enough to contain all of the data) will fix this problem.
--Bob Y. 15:57, 22 June 2012 (EDT)
Stack overflow
Finally, a segmentation fault can happen if GEOS-Chem uses up all of the available stack memory on your system. The stack memory is a special part of the memory where short-term variables get stored.
The compiler will typically place into the stack memory all local temporary variables, such as:
- variables that are local to a given subroutine
- variables that are NOT located within a COMMON block
- variables that are NOT declared with the SAVE attribute
- variables that are NOT declared as an ALLOCATABLE array
- variables that are NOT declared as a POINTER variable or array
Therefore, it is important to make sure that your computational environment is set up to use the maximum amount of stack memory. You can do this by placing the following line in your .cshrc file:
limit stacksize unlimited
or .bashrc file:
ulimit -s unlimited
If you encounter a SIGSEGV(174) message due to a stacksize memory error, you may see the following error text:
severe (174): SIGSEGV, possible program stack overflow occurred Program requirements exceed current stacksize resource limit.
--Bob Y. 15:57, 22 June 2012 (EDT)
forrtl: error (76): IOT trap signal
Xun Jiang wrote:
- We met the following error message
forrtl: severe (174): SIGSEGV, segmentation fault occurred Stack trace terminated abnormally. forrtl: error (76): IOT trap signal Note: The error appears after - RDSOIL: Reading Data/GEOS_2x2.5/soil_NOx_200203/climatprep2x25.dat ### MAIN: a DAILY DATA
- I have the following lines in .cshrc
setenv KMP_STACKSIZE 329033024 limit cputime unlimited limit datasize unlimited limit stacksize unlimited limit filesize unlimited limit memoryuse unlimited limit descriptors unlimited
- However, it still doesn't work. Any suggestion is really appreciated.
Bob Yantosca replied:
- I found this internet post which has an explanation:
Cause: The stack size for child threads are overflowing. The main stack size for the program is changed by the ulimit command (in Bash shell) or limit command (in C shell). However this environment variable does not set the size for the child thread stack size. Thus the child thread stack overflow. Solution: Set the environment variables to increase the child thread stack size. #for intel, using bash shell export KMP_STACKSIZE=500000000 # for intel, using csh or tcsh shell setenv KMP_STACKSIZE 500000000
- For more information, please see our wiki post on Resetting the stack size for Linux.
--Bob Y. 11:20, 26 June 2012 (EDT)
Segmentation fault encountered after TPCORE initialization
You may encounter a segmentation fault right after the following text is printed.
NASA-GSFC Tracer Transport Module successfully initialized
This error usually occurs when:
- You are running GEOS-Chem at sufficiently fine resolution, such as 2° x 2.5° or finer. (Many users have reported that this error does not occur at 4° x 5° resolution.)
- You are using a large number of advected tracers.
- Both #1 and #2
If you are using the Intel Fortran Compiler, the cause of this error can likely be traced to a known issue with the the glibc library. This will cause GEOS-Chem to think that it has used up all of the available memory, when in fact there is plenty of memory still available. However, you may also encounter this same error even if you have compiled GEOS-Chem with a different compiler.
You can usually correct this error by manually telling your system to use the maximum amount of stack memory when running GEOS-Chem. For detailed instructions, please see the following links:
- Setting stacksize for the Intel Fortran Compiler (aka "IFORT")
- Setting stacksize for the PGI Compiler
- Setting stacksize for the Sun Studio compiler
--Bob Y. 16:07, 14 December 2010 (EST)
Bad GEOS-4 A6 met data causing segmentation fault
Please see this post about bad GEOS-4 A6 met data causing a segmentation fault in GEOS-Chem simulations.
--Bob Y. 15:19, 16 February 2010 (EST)
The errors listed below, which occur infrequently, are related to invalid memory operations (especially with POINTER-based variables).
Bus Error
A bus error means that you are trying to reference memory that cannot possibly be there. The website StackOverflow.com has a definition of bus error and how it differs from a segmentation fault.
One cause of a bus error can be if you are trying to call a subroutine with the wrong number of arguments (i.e. usually too many arguments).
--Bob Y. 12:27, 19 October 2012 (EDT)
Dwarf subprogram entry error
The error message:
Dwarf subprogram entry L_ROUTINE-NAME__LINE-NUMBER__par_loop2_2_576 has high_pc < low_pc. This warning will not be repeated for other occurrences.
can occur when you try to use a pointer variable that is unassociated (i.e. that is not currently pointing to any other variable) from within an OpenMP parallel loop, where:
- ROUTINE-NAME is the name of the routine where the error occurred, and
- LINE-NUMBER is the line where the error occurred.
We recently discovered that this error can be caused if you have a pointer declaration such as this:
TYPE(Species), POINTER :: ThisSpc => NULL()
where the pointer ThisSpc is later used to point to another variable from within an OpenMP parallel loop. As it turns out, the above declaration statement will inadvertently cause pointer ThisSpc to be declared with the SAVE attribute. This can cause a segmentation fault, because all pointers used within an OpenMP parallel region must be created and destroyed on the same thread.
This type of problem can usually be fixed by removing the nullification from the declaration statement. In other words, you can rewrite the above line of code with:
TYPE(Species), POINTER :: ThisSpc . . . ThisSpc => NULL()
For more information, please see this article.
--Bob Yantosca (talk) 19:27, 29 April 2016 (UTC)
IFORT error: Relocation truncated to fit
Please see this wiki post on our Intel Fortran Compiler page which describes how to work around an Relocation truncated to fit error message.
--Bob Y. 10:46, 24 February 2012 (EST)
IFORT error: Out of memory asking for NNNNN
This is not a common error message, but it may occur if you are compiling a version of GEOS-Chem for a high-resolution horizontal grid, or with one of the available microphysics packages (i,e. APM or TOMAS). Please see this wiki post on our Intel Fortran Compiler page which describes this error in detail.
--Bob Y. 10:42, 26 July 2013 (EDT)
Memory error: "munmap_chunk: invalid pointer"
The following error is not common but can happen:
| Error | *** glibc detected *** ./geos: munmap_chunk(): invalid pointer: 0x00000000059aac30 *** |
|---|---|
| Reference | http://stackoverflow.com/questions/6199729/how-to-solve-munmap-chunk-invalid-pointer-error-in-c |
| Explanation | This happens when the pointer passed to (C-library language routine free(), which is called from Fortran routine NULLIFY()) is not valid or has been modified somehow. I don't really know the details here. The bottom line is that the pointer passed to free() must be the same as returned by (C-library routines) malloc(), realloc() and their friends.
The free() function frees the memory space pointed to by ptr, which must have been returned by a previous call to malloc(), calloc() or realloc(). Otherwise, or if free(ptr) has already been called before, undefined behavior occurs. If ptr is NULL, no operation is performed. GNU 2012-05-10 MALLOC(3) |
| Simpler explanation | This can happen if you are trying to deallocate or nullify a pointer variable that has already been deallocated or modified. |
--Bob Yantosca (talk) 21:32, 6 January 2017 (UTC)
Memory error: "free: invalid size"
The following error is not common but can happen:
| Error | *** Error in `./geos': free(): invalid size: 0x000000000662e090 *** |
|---|---|
| Reference | http://stackoverflow.com/questions/4729395/error-free-invalid-next-size-fast |
| Explanation | It means that you have a memory error. You may be trying to free a pointer that wasn't allocated (or delete an object that wasn't created) or you may be trying to nullify/delete such an object more than once. You may be overflowing a buffer or otherwise writing to memory to which you shouldn't be writing, causing heap corruption.
Any number of programming errors can cause this problem. You need to use a debugger, get a backtrace, and see what your program is doing when the error occurs. If that fails and you determine you have corrupted the heap at some previous point in time, you may be in for some painful debugging (it may not be too painful if the project is small enough that you can tackle it piece by piece). |
--Bob Yantosca (talk) 21:32, 6 January 2017 (UTC)
Memory error: "double free or corruption"
The following error is not common, but can occur under some circumstances:
| Error | *** glibc detected *** ./geos: double free or corruption (out): |
|---|---|
| Reference | http://stackoverflow.com/questions/2902064/how-to-track-down-a-double-free-or-corruption-error-in-c-with-gdb |
| Explanation | There are at least two possible situations:
For the first one I strongly suggest NULL-ing all deleted pointers. You have [some] options:
Three basic rules:
Combination of these three works quite well. |
--Bob Yantosca (talk) 21:32, 6 January 2017 (UTC)
Obsolete error messages
The following error messages occurred in older GEOS-Chem versions, prior to the implementation of HEMCO (in v10-01) and FlexChem (in v11-01).
Error caused by MEGAN biogenic emissions
This information only applies to GEOS-Chem v9-02 and prior versions. In GEOS-Chem v10-01 and higher versions, MEGAN biogenic emissions are handled by the HEMCO emissions component.
MEGAN keeps a 10-day running average of temperature, and therefore requires that the the met field files for the 10 days prior to the start of the GEOS-Chem simulation be present on disk.
You can solve this error in one of two ways:
- Make sure you have the previous 10 days (or better yet, the entire previous month!) of data prior to your GEOS-Chem simulation's starting date
- Start your GEOS-Chem simulation at a later date
Error caused by not recompiling cleanly
This error may also indicate that you have attempted to change the resolution of GEOS-Chem without recompiling cleanly. Some parts of the GEOS-Chem code (object files, module files) may still be compiled for a different grid resolution.
You can usually fix this doing a
make realclean
command before recompiling the code.
--Bob Y. 10:47, 15 November 2010 (EST)
I/O Error #29
This error indicates that GEOS-Chem cannot find the proper A3 met field file.
NOTE: Error #29 is specific to the IFORT compiler. If you are using a different compiler, then the I/O error number may differ.
--Bob Y. 15:30, 3 November 2010 (EDT)
Problem reading binary punch file
NOTE: Binary punch file I/O is being phased out of GEOS-Chem.
If you are having problems reading a binary punch file into GEOS-Chem, make sure that you have the correct endian setting in your makefile. These are:
- Intel Fortran compiler (IFORT): -convert big_endian
- PGI compiler: -byteswapio
- Sun Studio compiler: -xfilebyteorder=big16:%all
Most machines that use an Intel or AMD chipset are little-endian machines. A few of the older architectures (e.g. Cray, SGI Origin) are big-endian. Binary punch files are always big-endian (due to historical reasons), so you will need to tell your compiler to do the byte swapping manually.
The symptoms of such an error can be as follows:
Daewon Byun wrote:
- In the SUBROUTINE READ_BPCH2, It reads the FTI = CTM bin 02 fine, but then fails to read anything after. I dumped the IOUNIT and IO error code -- as you see TMP_TITLE is empty....
IUNIT, IOS, TMP_TITLE = 98 -1
- Then the program stops at the
IF ( IOS /= 0 ) THEN
PRINT*, 'open_bpch2_for_read:2'
STOP
ENDIF
- If I force to read further removing the "STOP", then I get (again, I tried to dump..)
MODELNAME, LONRES, LATRES, HALFPOLAR, CENTER180 = 0.000000 0.000000 0 0
--Bob Y. 09:57, 24 July 2008 (EDT)
Error when reading the "restart_gprod_aprod" file
NOTE: The GEOS-Chem SOA simulations no longer read the GPROD/APROD restart file.
Eric Leibensperger wrote:
- I am trying to run GEOS-Chem and have encountered and error. The log file gives me this:
=============================================================================== GEOS-CHEM ERROR: No matches found for file restart_gprod_aprod.2001070100! STOP at READ_BPCH2 (bpch2_mod.f)! ===============================================================================
- I have the aerosol restart file (with the same name) in my ~/testrun/runs/run.v7-04-12/ folder. Is it looking for it elsewhere? I get an additional message in the log.error file, but I think that it is possibily the result of not being able to find the file above:
****** FORTRAN RUN-TIME SYSTEM ****** Error 1183: deallocating an unallocated allocatable array Location: the DEALLOCATE statement at line 4933 of "carbon_mod.f" Abort
- Any thoughts would be appreciated. Sorry to bother you with this!
- Eric
Philippe Le Sager replied:
- You must rewrite your restart_gprod_aprod.YYYYMMDDHH so that the date in the filename is the same as the one in the datablock header.
- I wrote a routine to do that: ~phs/IDL/dvpt/various_rewrite/rewrite_agprod.pro
- -Philippe
NOTE: The file rewrite_agprod.pro will be released in the next GAMAP version.
--Bmy 15:59, 9 May 2008 (EDT)
For GEOS-Chem v8-03-01 and higher
In GEOS-Chem v8-03-01 and higher, the restart_gprod_aprod.YYYYMMDDhh file has been renamed to soaprod.YYYYMMDDhh. The above-described error can occur with the soaprod.YYYYMMDDhh file if the date in the file does not match the starting date of your simulation.
--Bob Y. 13:10, 4 November 2010 (EDT)
File ann_mean_trop.geos5.* not found
The fixed annual-mean tropopause is only used for simulations with GEOS-4, which is being phased out.
If you are running a GEOS-5 simulation and get an error that says that GEOS-Chem cannot locate the ann_mean_trop.geos5.2x25 or ann_mean_trop.geos5.4x5 file, then make sure that the following option is set in your input.geos file.
Use variable tropopause?: T
Starting in version GEOS-Chem v7-04-12, GEOS-Chem can now use a variable tropopause (i.e. chemistry is done up to the location of the actual tropopause as diagnosed from the met fields at any given timestep). You cannot use the annual mean tropopause for GEOS-5.
--Bob Y. 15:18, 7 July 2008 (EDT)
Problem reading GEOS-4 TROPP files
The fixed annual-mean tropopause is only used for simulations with GEOS-4, which is being phased out.
Please see this wiki post for more information about a common problem that can occur if you using GEOS-4 meteorology with the dynamic tropopause.
--Bob Yantosca (talk) 21:02, 6 January 2017 (UTC)
OPEN failure
This code has since been removed from GEOS-Chem. All emissions are now computed via HEMCO.
If you are reading data from a binary punch file, and encounter this type of error:
- ANTHRO_CARB_TBOND: Reading /as/group/geos/data/GEOS_1x1/historical_emissions_201203/BCOC/BCOC_anthsrce.2000.geos.1x1 Error opening filename= ^@^@^@^@^@^@^@^@^@^@^@^@^@^@^@^@^@^@^@^@^@^@^@^@^@^@^@^@^@^@^@^@^@^@^@^@^@^@^@^@^@^@^@^@ <AA><A6>B^@^@^@^@^@^@^@^@^@^@^@^@^@^@^@ =============================================================================== GEOS-CHEM I/O ERROR 30 in file unit 65 Encountered at routine:location open_bpch2_for_read:1 Error 30: OPEN failure ===============================================================================
Then recompile your code with the BOUNDS=yes option. Chances are this is the side-effect of an array out-of-bounds error caused by your array being too small to hold the data that is being read in from disk. Compiling with BOUNDS=yes and re-running should help you locate the problem, and you should get an error message such as:
forrtl: severe (408): fort: (2): Subscript #1 of the array ARRAY has value 73 which is greater than the upper bound of 72
which will point you to the specific array where the problem is occurring.
--Bob Yantosca (talk) 21:03, 6 January 2017 (UTC)
SMVGEAR "Too many decreases of YFAC" error
SMVGEAR has now been removed from GEOS-Chem v11-01 and higher versions. The following information is obsolete, but we shall leave it here for reference.
If you are performing a full-chemistry simulation with the SMVGEAR solver, you may experience this error. This will cause error output (similar to what is listed below) to your log file.
- PHYSPROC: Trop chemistry at 2006/01/01 00:00 SMVGEAR: DELT= 8.68E-16 TOO LOW DEC YFAC. KBLK, KTLOOP, NCS, TIME, TIMREMAIN, YFAC, EPS = 422 24 1 1.800E+03 5.422E+02 1.000E+00 1.000E-01 SMVGEAR: DELT= 5.91E-16 TOO LOW DEC YFAC. KBLK, KTLOOP, NCS, TIME, TIMREMAIN, YFAC, EPS = 428 24 1 1.800E+03 1.069E+03 1.000E+00 1.000E-01 SMVGEAR: DELT= 6.14E-16 TOO LOW DEC YFAC. KBLK, KTLOOP, NCS, TIME, TIMREMAIN, YFAC, EPS = 434 24 1 1.800E+03 1.214E+03 1.000E+00 1.000E-01 SMVGEAR: DELT= 6.79E-16 TOO LOW DEC YFAC. KBLK, KTLOOP, NCS, TIME, TIMREMAIN, YFAC, EPS = 422 24 1 1.800E+03 6.323E+02 1.000E-02 1.000E-01 ... etc ... SMVGEAR: TOO MANY DECREASES OF YFAC M1,M2,K,ERR = 72 25 1 4.3954E-04 M1,M2,K,ERR = 72 26 1 4.6620E-04 M1,M2,K,ERR = 72 27 1 1.3593E-04 M1,M2,K,ERR = 72 28 1 1.0686E-04 M1,M2,K,ERR = 72 29 1 1.6856E-04 ... etc ... CONC WHEN STOP = 1 1 DRYCH2O 0.00E+00 0.00E+00 CONC WHEN STOP = 2 1 DRYH2O2 0.00E+00 0.00E+00 CONC WHEN STOP = 3 1 DRYHNO3 0.00E+00 0.00E+00 CONC WHEN STOP = 4 1 DRYN2O5 0.00E+00 0.00E+00 ... etc ...
Each of the lines starting with
SMVGEAR: DELT=_______ TOO LOW DEC YFAC
is a warning that the SMVGEAR gear solver could not converge to a solution in a particular grid box. In this case, SMVGEAR does not stop with an error, but instead reduces the internal solver timestep and tries to solve the chemistry in that grid box once again.
But if SMVGEAR cannot converge to a solution in the grid box after reducing the internal timestep a certain number of times, it will give up on solving the chemistry and halt the run with an error. This is indicated by the lines:
SMVGEAR: TOO MANY DECREASES OF YFAC M1,M2,K,ERR = 72 25 1 4.3954E-04 ... etc ...
SMVGEAR will then proceed to print out the concentration of each chemical species in the matrix, which is indicated by lines such as these:
CONC WHEN STOP = 1 1 DRYCH2O 0.00E+00 0.00E+00
There can be several causes for this error:
- If you are splitting up your simulation into separate stages (i.e. so that you can fit each year’s run into your queue’s time limits), then it is important to turn on the chemical species restart file in the input.geos options file. This ensures that the concentrations of the chemical species will be preserved from one GEOS-Chem run stage to the next. Otherwise, SMVGEAR will re-initialize the chemical species to zero at the start of the next GEOS-Chem run stage. This may cause the SMVGEAR solver not to be able to converge to a solution.
- An error may occur in a different area of GEOS-Chem (such as wet deposition) that causes bad values (e.g NaN, negative numbers, Infinity) to be written to array of tracer concentrations or chemical species concentrations. These bad values can then cause the SMVGEAR solver to halt.
If you should encounter this error, here are a few things that you can try:
- Check your restart files for bad values.
- If you are not using an "out-of-the-box" version of GEOS-Chem, then check the areas of the code that you have modified for potential errors.
- Recompile GEOS-Chem from scratch with the BOUNDS=yes option turned on. This will flag any array-out-of-bounds errors.
- Run a series of short (1-2 day simulations) in which you successively turn off one other operation (i.e. transport, convection, wetdep, PBL, etc) at a time. This may help you to localize the source of the error.
- Run a short GEOS-Chem simulation with OpenMP parallelization turned off (i.e. use the OMP=no option). A parallelization error may be corrupting values in arrays that are needed by SMVGEAR.
- If you have access to a debugger like IDB or TotalView, recompile GEOS-Chem with DEBUG=yes and then run GEOS-Chem in the debugger.
--Bob Y. 10:39, 26 July 2013 (EDT)
JXTRA error in FAST-JX photolysis
This issue was resolved in GEOS-Chem v11-01d (approved 12 Dec 2015).
SMVGEAR has now been removed from v11-01g and higher versions. The following information is obsolete, but we shall leave it here for reference.
Those of you who are using the GEOS-Chem v10-01 public release code may encounter this error.
Doug Finch wrote:
A few of us here at Edinburgh are running v10 of the model and coming across the same error:
SMVGEAR: TOO MANY DECREASES OF YFAC
This happens after lots of write outs of this:
N_/L2_/L2-cutoff JXTRA: 601 96 0.00
This error seems to be occurring a fairly random times during runs of different resolutions (for example 25th of October 2014 at 2x25 deg). These are all full chemistry runs.
We've been trying to get to the bottom of this but we're not sure why it is happening. We have a theory it is to do with the met fields but this isn't really based on much. Do you have any idea why this could be happening and how we can solve it?
Bob Yantosca replied:
Thanks for writing. It may be that you need to add this fix into your code. We officially added this fix in GeosCore/convection_mod.F in GEOS-Chem v11-01d.
Without this fix, you could have extremely high concentrations of tracer (when using GEOS-FP or MERRA met). This could in turn cause the FAST-JX solver to choke with the error that you reported. If the aerosol concentration in a grid box gets too high, then FAST-JX doesn't know how to deal with the photolysis and just dies w/ an error.
--Bob Yantosca (talk) 21:19, 14 March 2016 (UTC)
Too many levels in photolysis code
FAST-J was replaced by FAST-JX v7.0 in GEOS-Chem v10-01 (approved 15 Jun 2015).
Please see this discussion about the "Too many levels in photolysis code" error that can sometimes happen in the FAST-J photolysis code.
--Bob Y. 11:09, 12 January 2010 (EST)
Error computing F_OF_PBL
The define.h include file was removed from v9-02 and higher versions This is no longer an issue.
If you should encounter this error (which occurs in routine COMPUTE_PBL_HEIGHT from pbl_mix_mod.f)
bad sum at: 1 70 -1.00000000000000
bad sum at: 1 81 -1.00000000000000
===============================================================================
GEOS-CHEM ERROR: Error in computing F_OF_PBL!
STOP at COMPUTE_PBL_HEIGHT ("pbl_mix_mod.f")
===============================================================================
===============================================================================
GEOS-CHEM ERROR: Error in computing F_OF_PBL!
STOP at COMPUTE_PBL_HEIGHT ("pbl_mix_mod.f")
===============================================================================
- CLEANUP: deallocating arrays now...
bad sum at: 1 48 -1.00000000000000
bad sum at: 1 59 -1.00000000000000
bad sum at: 73 46 -1.00000000000000
you should check to see if your code was not recompiled cleanly. Often this is a result of not doing a
make realclean
before trying to switch from 2° x 2.5° to 4° x 5° or vice versa in the define.h header file. Often this type of error can be deduced by looking at the GEOS-Chem log file. When this error occurs the resolution indicated at the top of the GEOS-Chem log file, e.g.:
************* S T A R T I N G 2 x 2.5 G E O S--C H E M *************
will not match the longitudes and latitudes as reported a little further down in the log file:
Grid box longitude centers [degrees]: -180.000 -175.000 -170.000 -165.000 -160.000 -155.000 -150.000 -145.000 ... Grid box latitude centers [degrees]: -89.000 -86.000 -82.000 -78.000 -74.000 -70.000 -66.000 -62.000 ...
--Bob Y. 12:13, 25 October 2010 (EDT)
LISOPOH error
SMVGEAR has now been removed from GEOS-Chem v11-01 and higher versions. The following information is obsolete, but we shall leave it here for reference.
If you are running the SOA simulation and encounter this error:
=============================================================================== GEOS-CHEM ERROR: LISOPOH needs to be defined for SOA! STOP at chemdr.f ===============================================================================
Then this means that the species LISOPOH is not set to an active species in the chemical mechanism. LISOPOH needs to be a turned off for the standard 43-tracer simulation, but turned on the SOA simulation.
To fix the error, change this line in your globchem.dat file from:
D LISOPOH 1.00 1.000E-20 1.000E-20 1.000E-20 1.000E-20
to
A LISOPOH 1.00 1.000E-20 1.000E-20 1.000E-20 1.000E-20
and then re-run GEOS-Chem.
--Bob Yantosca (talk) 21:06, 6 January 2017 (UTC)
Fatal error in IFORT
NOTE: CMN_DIAG.h (and most other include files) were converted into Fortran-90 modules in GEOS-Chem v9-02. This information is now obsolete.
The following error, which resulted on the Altix platform using Intel "ifort" v9.1 compiler:
ifort: error: /opt/intel/fc/9.0/bin/fpp: core dumped ifort: error: Fatal error in /opt/intel/fc/9.0/bin/fpp, terminated by unknown signal(139) make: *** [transport_mod.o] Error 1
was caused by an omitted " in an #include declaration, i.e.
# include "CMN_DIAG
Adding the closing " fixed the problem.
A3 met fields not found
NOTE: This error only applies to the GEOS-4, GEOS-5, MERRA, and GCAP met fields, which are stored in binary format. GEOS-FP and MERRA-2 met fields are read from netCDF format files.
If routine OPEN_A3_FIELDS (in a3_read_mod.f) returns a "file not found" error shortly after the start of your GEOS-Chem simulation, i.e.:
$$ Finished Reading Linoz Data $$
===============================================================================
GEOS-CHEM I/O ERROR 29 in file unit 73
Encountered at routine:location open_a3_fields:1
Error 29: File not found
===============================================================================
- CLEANUP: deallocating arrays now...